Lyngbya sp. (strain PCC 8106) (Lyngbya aestuarii (strain CCY9616))
Taxonomy: cellular organisms; Bacteria; Terrabacteria group; Cyanobacteria/Melainabacteria group; Cyanobacteria; Oscillatoriophycideae; Oscillatoriales; Lyngbya
Average proteome isoelectric point is 6.22
Get precalculated fractions of proteins
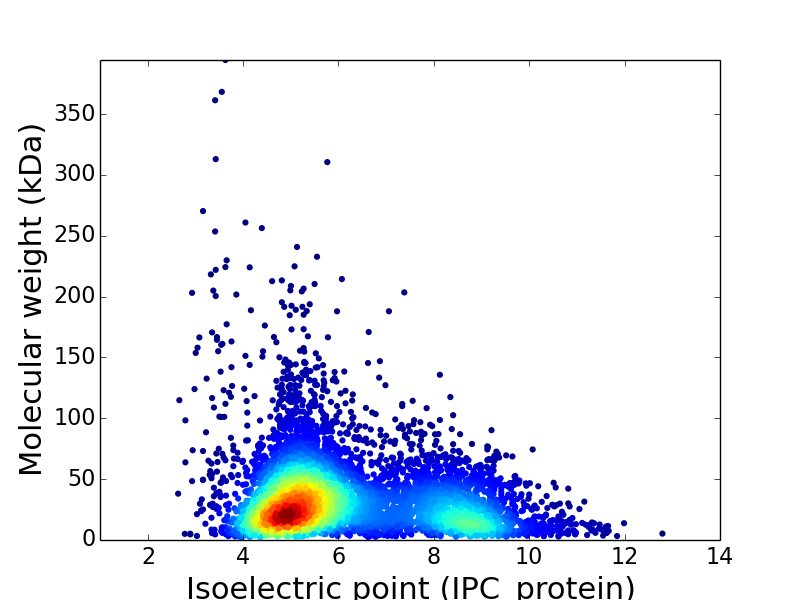
Virtual 2D-PAGE plot for 6,110 proteins (isoelectric point calculated using IPC_method)
Get csv file with sequences according given criteria:
* You can choose from 18 different methods for calculating isoelectric point
Protein with the lowest isoelectric point:
>tr|A0YIP8|A0YIP8_LYNSP Endonuclease/exonuclease/phosphatase
MTTIINFDELAGNTVVSDQYSGVTFSTDPGSVVAIPIAALFGTSAPNIIGTFENGGLNGNLTLDVAFESPVNNFSFFTIGDNTNGTAALIDVYTADGLTTTVDLITDGNIFTPELVDLSSFTNVTRINVNTITDVGGIGYDDFAFEFAPLNIAPVAVDDSATAAQNSSQTILAADLLANDTDEDGDPLTL
TEVSNPLNGTVALDANGDVVFTPTSGFSGTASFEYTVSDGTDTDIGSVAVEVGGMFNGGNGKDTLIGTGGNDWMSGGNGADELYGGDGDDTLGGDGSDNGPDLLDGGLGNDILTGGSGPDVFVLAPGNGTDTITDFDTPDSIGLSGGIGFGDLSFSGSNIILTSTSEVLATLVSVDATSLTANDFVGI
MTTIINFDELAGNTVVSDQYSGVTFSTDPGSVVAIPIAALFGTSAPNIIGTFENGGLNGNLTLDVAFESPVNNFSFFTIGDNTNGTAALIDVYTADGLTTTVDLITDGNIFTPELVDLSSFTNVTRINVNTITDVGGIGYDDFAFEFAPLNIAPVAVDDSATAAQNSSQTILAADLLANDTDEDGDPLTL
TEVSNPLNGTVALDANGDVVFTPTSGFSGTASFEYTVSDGTDTDIGSVAVEVGGMFNGGNGKDTLIGTGGNDWMSGGNGADELYGGDGDDTLGGDGSDNGPDLLDGGLGNDILTGGSGPDVFVLAPGNGTDTITDFDTPDSIGLSGGIGFGDLSFSGSNIILTSTSEVLATLVSVDATSLTANDFVGI
Molecular weight: 38.08 kDa
Isoelectric point according different methods:
Isoelectric point according different methods:
Protein with the highest isoelectric point:
>tr|A0YYM2|A0YYM2_LYNSP 50S ribosomal protein L34
MAQQTLKGTNRKKKRVSGFRVRMRTKNGQSVIRSRRKKGRARLSV
MAQQTLKGTNRKKKRVSGFRVRMRTKNGQSVIRSRRKKGRARLSV
Molecular weight: 5.26 kDa
Isoelectric point according different methods:
Isoelectric point according different methods:
General Statistics
Number of major isoforms |
Number of additional isoforms |
Number of all proteins |
Number of amino acids |
Min. Seq. Length |
Max. Seq. Length |
Avg. Seq. Length |
Avg. Mol. Weight |
|---|---|---|---|---|---|---|---|
1,945,103 |
20 |
3,820 |
321.3 |
36.0 kDa |
Amino acid frequency
Ala |
Cys |
Asp |
Glu |
Phe |
Gly |
His |
Ile |
Lys |
Leu |
Met |
Asn |
Gln |
Pro |
Arg |
Ser |
Thr |
Val |
Trp |
Tyr |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
7.05 |
1.01 |
5.1 |
6.76 |
4.09 |
6.54 |
1.8 |
6.92 |
4.89 |
10.76 |
1.78 |
4.67 |
5.72 |
4.68 |
4.84 |
6.74 |
5.81 |
6.38 |
1.43 |
3.05 |
For dipeptide frequency statistics click here
Proteome-pI is available under Creative Commons Attribution-NoDerivs license, for more details see here
| Reference: Kozlowski LP. Proteome-pI: proteome isoelectric point database. Nucleic Acids Res. 2016. doi: 10.1093/nar/gkw978 |
Contact: Lukasz P. Kozlowski |

